Values
Details...
Projects
The Harms Lab studies protein evolution and biophysics using experimental and computational methods.
Current Members

Mike
Harms
Principal Investigator
Favorite part of job: trainees waving graphs in office.

Jon
Muyskens
Lab Manager
Developmental biologist turned evolutionary biochemist.

Michael
Shavlik
PhD student
Experimental studies of "might-have-been" evolutionary trajectories

Kona
Orlandi
PhD student
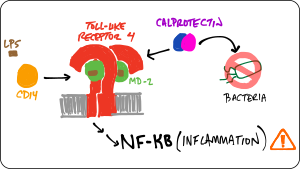
Early evolution of TLR4

Lauren
Lehmann
PhD student
How does S100A9 interact with TLR4?

Sophia
Phillips
PhD student
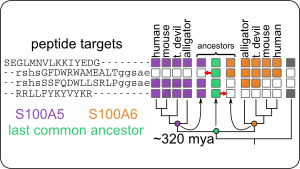
Evolution of S100A9 proinflammatory activity

Jose
Sanchez-Borbon
PhD student
How did the CD14 LPS binding site evolve?
Previous Members
(in reverse order of when each left)

Luis
Perez
PhD student
Studies of protein sequence space

JJ
Yin
Undergrad researcher
Evolution of S100A9 proinflammatory activity

Anneliese
Morrison
PhD, 2021
Research scientist with Absci

Hannah
Murawsky
Undergrad, 2021
Biochemistry (honors thesis), med student at UNLV

Daria
Wonderlick
Undergrad, 2021
Biochemistry (honors thesis), PhD student at Stanford

Joseph
Harman
PhD, 2020
Postdoc with David Baker, U Washington

Patrick
Connor
Undergrad
Biochemistry, med student at Iowa

Gus
Warren
Undergrad
Biochemistry, research tech at OHSU

Nick
Frantz
Masters, 2020
Biochemistry

Jeremy
Anderson
Postdoc
Data scientist, Travelers Insurance

Genevieve
Dorrell
Undergrad
Marine Biology

Leander
Goldbach
Visiting Masters Student

Andrea
Loes
PhD, 2018
Lab manager, Bloom Lab, Fred Hutch

Zach
Sailer
PhD, 2018
Software developer, Apple

Caitlyn
Wong
Undergrad
Biochemistry (honors thesis), med student at OHSU

Luke
Wheeler
PhD, 2017
Postdoc with Stacey Smith, UC Boulder

Ran
Shi
Undergrad
Biochemistry, PhD student at U. Georgia

Bram
Rickett
Undergrad
Biochemistry (honors thesis)

Hiranmayi
Duvvuri
Undergrad
Biochemistry

Maureen
Heaphy
Undergrad
Marine Biology

Jaclyn
Smith
Undergrad
Biology
Publications
An experimental demonstration of ensemble epistasis in the lac repressor
bioRxiv
Disentangling contact and ensemble epistasis in a riboswitch
Biophysical Journal
Higher-order interactions shape microbial interactions as microbial community complexity increases
Scientific Reports 12: 22640
Topiary: pruning the manual labor from ancestral sequence reconstruction
Protein Science
Evolution avoids a pathological stabilizing interaction in the immune protein S100A9
PNAS 119 (41) e2208029119
Ensemble epistasis: thermodynamic origins of non-additivity between mutations
Genetics 219(1):iyab105
A conserved folding nucleus sculpts the free energy landscape of bacterial and archaeal orthologs from a divergent TIM barrel family
PNAS 118(17):e2019571118
Evolutionary conservation of structural and functional coupling between the BRM AT-hook and bromodomain
Journal of Molecular Biology 166845
Exploring the evolutionary history of kinetic stability in the alpha-lytic protease family
Biochemistry 60(3):170–181
Were ancestral proteins less specific?
Molecular Biology and Evolution msab019
Identification and characterization of zebrafish Tlr4 co-receptor Md-2
Journal of Immunology 206(5):1046-1057
Learning Peptide Recognition Rules for a Low Specificity Protein
Protein Science 29:2259 –2273
Inferring a complete genotype-phenotype map from a small number of measured phenotypes
PLOS Computational Biology e1008243
Evolution of multifunctionality through a pleiotropic substitution in the innate immune protein S100A9
eLife 9:e54100
Zinc-independent activation of Toll-like receptor 4 by S100A9
bioRxiv
Skeletal geometry and nice transitions restore organ size and shape during zebrafish fin regeneration
bioRxiv
Tracing the evolution of novel features of human Toll-like receptor 4
Protein Science 28(7):1350-1358
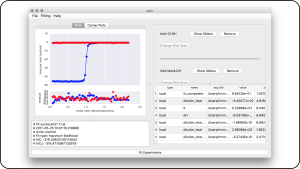
pytc: open source python software for global analyses of isothermal titration calorimetry data
Biochemistry 57(18):2578–2583
Coevolution of the Toll-Like Receptor 4 complex with calgranulins and lipopolysaccharide
Frontiers in Immunology
Conservation of specificity in two low-specificity proteins
Biochemistry 57(5):684-695
Enzymes emerge by upcycling [News & Views]
Nature Chemical Biology 14:526-527
Ca2+ and Cu2+ bind independently to human S100A5
BMC Biophysics 10:8
Molecular ensembles make evolution unpredictable
PNAS 114(45):11938-11943
High-order epistasis shapes evolutionary trajectories
PLOS Computational Biology
Detecting high-order epistasis in nonlinear genotype-phenotype maps
Genetics 205(3):1079-1088
Robustness of reconstructed ancestral protein functions to statistical uncertainty
Molecular Biology and Evolution 34 (2):247-261
Multiple evolutionary origins of ubiquitous Cu2+ and Zn2+ binding in the S100 protein family
PLOS ONE 11(10): e0164740
Following the folding trajectory of RNases H over evolutionary time: a trend towards kinetic stability
PNAS 113(46):13045-13050
Ancient protein thermostability and specificity [Invited Review]
Current Opinions in Structural Biology 38:37-43
Thermodynamic system drift in the evolution of protein thermostability
PLOS Biology 12(11):e1001994
Biophysical basis for historical contingency in glucocorticoid receptor evolution
Nature 512:203-207
Biophysical mechanisms for large-effect mutations in the evolution of steroid hormone receptors
PNAS 110(28):11475-11480
Evolutionary biochemistry: Revealing the historical and physical causes of protein function
Nature Reviews Genetics 14(8):559-571
Resurrection of an Urbilaterian U1A/U2B''/SNF protein
Journal of Molecular Biology 425(20):3846–3862
Evolution of minimal specificity in the steroid receptors
PLOS Genetics 8(11):e1003072
Arginine is ionized at internal positions in a protein
PNAS 108(47):18954-9
Protein evolution by molecular tinkering: diversification of the nuclear receptor superfamily from a ligand-dependent ancestor
PLOS Biology 8(10):e1000497
Analyzing protein structure and function using ancestral gene reconstruction
Current Opinions in Structural Biology 20(3):360-366
The pKa values of acidic and basic residues buried at the same internal location in a protein are governed by different factors
Journal of Molecular Biology 389:34-47
A buried lysine that titrates with a normal pKa: Role of conformational flexibility at the protein–water interface as a determinant of pKa values
Protein Science 17:833-845
How sequence defines structure: a crystallographic map of DNA structure and conformation
PNAS 102(20):7157-7162
Laser light-scattering evidence for an altered association of βB1-crystallin deamidated in the connecting peptide
Protein Science 13:678-686
Location
Location on campus
Mike's office: Willamette 340A
Lab: Willamette 340, 342
Address
Harms Lab
Institute for Molecular Biology
1229 University of Oregon
Eugene, OR 97403-1229
Phone
541-346-9003